Chinese Scientists Determine Structural Basis for Inhibition of the RNA-Dependent RNA Polymerase from SARS-CoV-2 by Remdesivir
A team of Chinese scientists has reported the high-resolution cryo-EM structure of RNA and remdesivir bound RNA-Dependent RNA Polymerase(RdRp) complex from SARS-CoV-2.
The research, published online in Science on May 1, was conducted by Prof. XU Huaqiang and Prof. XU Yechun from the Shanghai Institute of Materia Medica (SIMM) of the Chinese Academy of Sciences (CAS), Prof. ZHANG Yan from the Zhejiang University School of Medicine, Prof. ZHANG Shuyang from Peking Union Medical College and Chinese Academy of Medical Sciences, and their collaborators.
Since the outbreak of COVID-19, it has spread around the world, causing a huge disaster to all mankind. Many countries around the world still facing huge challenges to fight SARS-CoV-2, and it is very urgent to find an effective treatment.
The SARS-CoV-2 is a positive-strand RNA virus that mainly infects human cells through the mucosal system. The massive replication of the virus requires the rapid synthesis of its genetic RNA. This process is mediated by a multi-subunit replication transcription complex composed of multiple non-structural proteins (nsp) of the virus. The core element is the RdRp complex, which is the core component of coronavirus replication. Numerous nucleoside drugs currently under clinical testing, including Remdesivir, targeting RdRp. Remdesivir is a prodrug form that will be metabolized into the active product in the form of triphosphate in the body to cause the termination of the viral RNA chain extensio.
In order to elaborate the fine anti-viral mechanism of nucleoside drugs, after 46 days of hard work, the team successfully reported the cryo-EM structure of the SARS-CoV-2 RdRp either in the apo form at 2.8 ? resolution or in complex with a template-primer RNA and Remdesivir at 2.5 ? resolution.The overall conformation of the complex structure is very similar to that of the apo form, only a certain conformational change in some subdomains.
Comprehensive analysis shows that the SARS-CoV-2 RdRp complex is a very efficient enzyme. During RNA extension, the conformational change is small, which also explains the high contagious ability of the SARS-CoV-2; RdRp complex recognizes RNA but not DNA , in a way of sequence independent binding; the complex structure explains how Remdesivir enters the replication active site and covalently inserts into the viral genome, thereby inhibiting virus replication; The residues involved in RNA binding as well as residues comprising the catalytic active site are highly conserved among most RNA viruses including polio virus, hepatitis C virus, and influenza virus, etc. , showing the conservative mechanism of RdRp complex during gene replication, and suggesting it may be possible to develop broad spectrum antiviral inhibitors The structures provide a structural basis for the urgent development of potential antiviral drugs, helps to study potential binding patterns, and provides theoretical support for the design of more powerful, efficient and specific anti-SARS-CoV-2 drugs.
There are currently no specific drugs for COVID-19. Because of the low homology to host proteins, enzymes that are critical to the virus life cycle are excellent antiviral drug targets. Among them, the RdRp complex is the main target of many nucleoside drugs. This study reports the structure of the SARS-CoV-2 RdRp complex in the apo form and in the complex with a template-primer RNA and the active form of Remdesivir. These structures reveal how the template primer RNA is recognized, and the mechanism by which Remdesivir inhibits strand extension. Structure comparison and sequence alignment suggest that the mode of substrate RNA recognition and Remdesivir inhibition of RdRp is highly conserved in diverse RNA viruses, providing a basis for designing broad spectrum antiviral drugs based on nucleotide analogs. In addition, the structures also provide a solid template for modeling and modifying nucleotide drugs. In conclusion, these results provide a strong basis for designing more effective inhibitors to combat the malignant infection of the SARS-CoV-2.
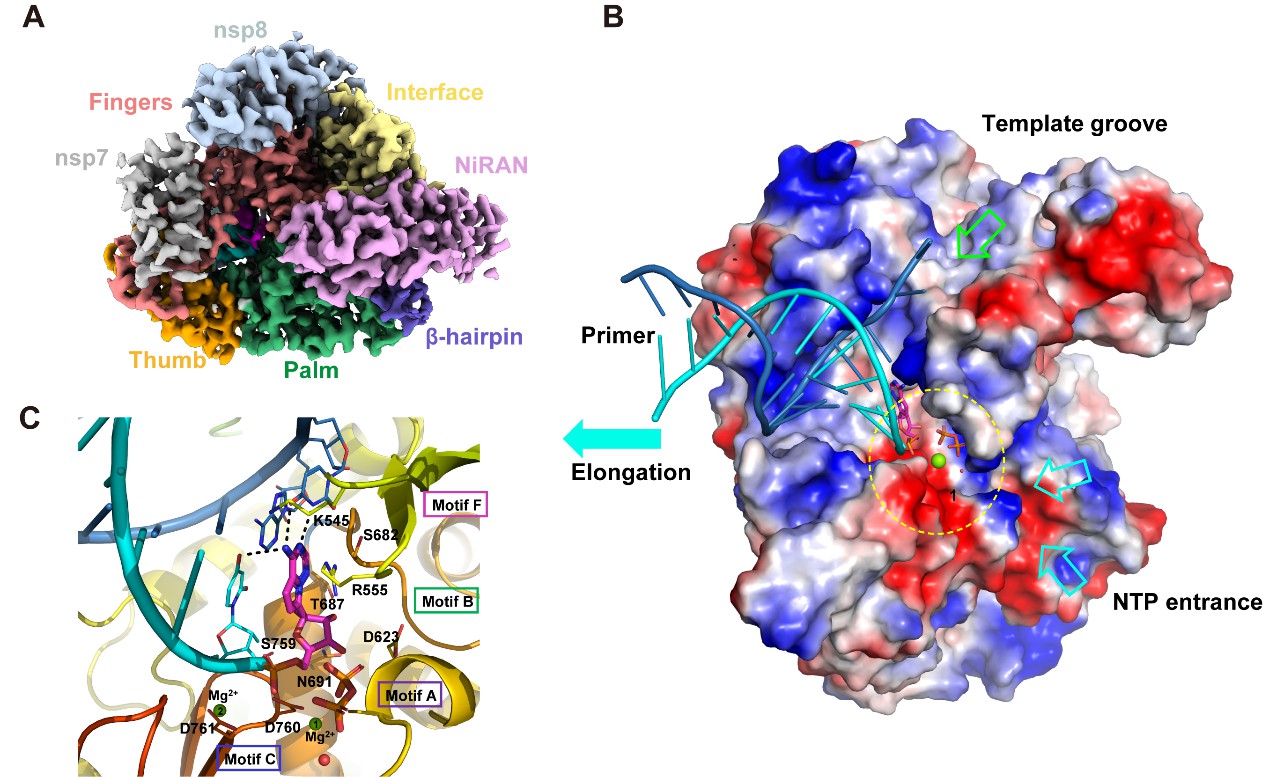
The cryo-EM structure of the RNA and Remdesivir bound RdRp complex from SARS-CoV-2.(A), EM map of the complex structure; (B) Surface view of the RdRp active site with the electrostatic potential; (C) A close view of the RdRp active site, showing the covalently bound RMP, pyrophosphate, and magnesium ions. Key residues and bases that interact with Remdesivir are shown.(Image by SIMM)
The article linker:https://science.sciencemag.org/lookup/doi/10.1126/science.abc1560
Contact: WANG Xiaocheng, wxc@simm.ac.cn




