New Tools Decode the Function of a Ketone-Driven Protein Modification
Protein post-translational modifications (PTMs) expand the functional diversity of the proteome well beyond what is encoded in the genome. By regulating protein activity, localization, stability, and molecular interactions, PTMs participate in virtually all cellular processes. One recently discovered PTM is lysine acetoacetylation (Kacac), driven by acetoacetate — a ketone body produced during fasting and fat metabolism. Since its discovery in 2023, Kacac has been detected on both histone and non-histone proteins in mammals. Yet the full landscape of Kacac substrates and its biological roles remained largely unknown, mainly due to the lack of sensitive and specific detection tools.
In a study published in Science Advances, a research team led by HUANG He at the Shanghai Institute of Materia Medica (SIMM), Chinese Academy of Sciences, and first author SONG Xiaohan, an assistant researcher at the Hangzhou Institute for Advanced Study, University of Chinese Academy of Sciences (UCAS), built an integrated platform combining chemical probe labeling with genetic code expansion, addressing this critical gap.
First, they designed a novel chemical probe called Aca-Bio. This probe exploits a unique feature of Kacac: its ketone carbonyl group. Aca-Bio forms a pH-responsive, reversible oxime bond with Kacac side chains. This allowed selective labeling, enrichment, and controlled release of Kacac-modified peptides from complex biological samples. Using this chemoproteomic strategy on liver tissue from fasted mice, the researchers identified 260 Kacac sites across 125 proteins. This substantially expanded the known Kacac substrate repertoire. The modified proteins were significantly enriched in core metabolic pathways, including the tricarboxylic acid (TCA) cycle and ketogenesis.
To move from discovery to function, the team developed a genetic code expansion system. This system enabled site-specific incorporation of Kacac at defined positions in mammalian cells. They applied it to 3-hydroxy-3-methylglutaryl-CoA synthase 2 (HMGCS2), a key enzyme in ketone body biosynthesis. Kacac at lysine 310 (K310) of HMGCS2 was found to be highly abundant. Structural analysis indicated that this modification reduces substrate binding through steric hindrance, leading to a marked decrease in enzymatic activity. This suggests Kacac may act as a “molecular brake” in metabolic pathways, participating in negative feedback regulation of ketogenesis.
This work provides a complete methodological pipeline from global Kacac site mapping to site-specific functional characterization. The tools developed here open new avenues for studying how metabolite-driven PTMs shape physiological and pathological processes. The findings also raise the possibility that Kacac modulates metabolic flux during fasting and metabolic stress, with potential implications for diseases such as diabetes and metabolic syndrome.
The study was supported by the Strategic Priority Research Program of the Chinese Academy of Sciences, the Research Funds of Hangzhou Institute for Advanced Study, and the National Natural Science Foundation of China.
DOI: 10.1126/sciadv.aeb5106
Link:https://www.science.org/doi/10.1126/sciadv.aeb5106
Keywords: lysine acetoacetylation; chemoproteomic profiling; genetic code expansion
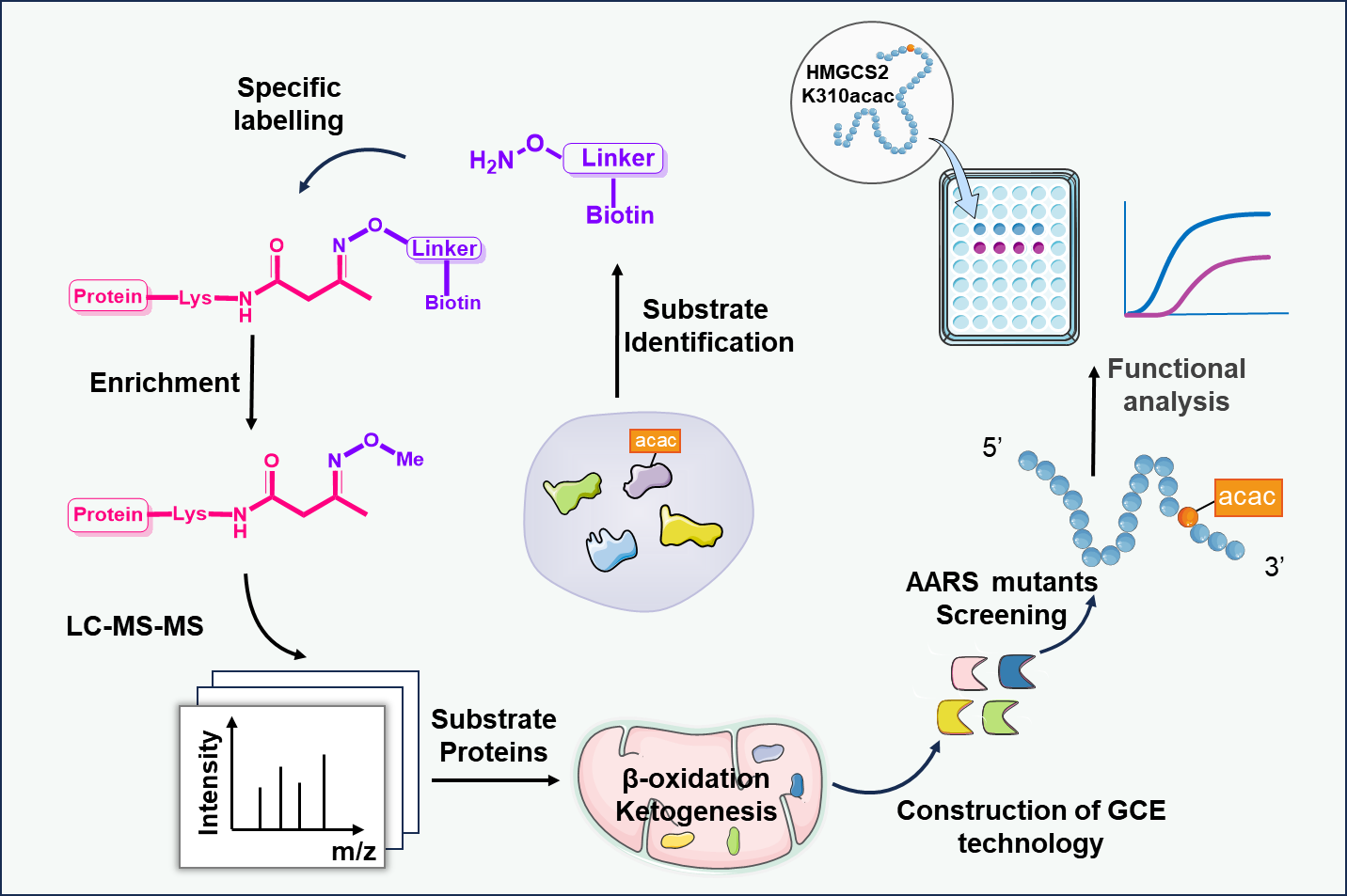
Schematic of the Aca-Bio probe strategy for Kacac enrichment and site-specific functional analysis. (Image by HUANG He’s lab)
Contact:
DIAO Wentong
Shanghai Institute of Materia Medica
E-mail: diaowentong@simm.ac.cn




