Researchers Revealed Assembly Program and Activation Mechanism of K63-polyUb Mediated MDA5-MAVS Antiviral Signal Pathway
MDA5 is an allogeneic RNA monitoring protein in cells and an important member of RIG-I-like receptor family (RLRs). MDA5, as an important barrier of natural immunity, is mainly involved in regulating the immune response caused by RNA viruses such as coronavirus, small RNA virus, flavivirus and hepatitis D virus. The RLRs family has three members: RIG-I, MDA5 and LGP2. The N-terminal of RIG-I and MDA5 have tandem CARDs domains. CARDs belong to the death domain superfamily, which can recruit MAVS through CARD-CARD homotypic interaction and finally promote the activation of type I interferon (IFN) pathway.
Ubiquitination is an important post-translational modification in eukaryotes. Ubiquitin can be modified by homo ubiquitin molecules and assembled into polyubiquitin chains, including M1, K11, K27, K48 and K63. Among them, K63-linked polyubiquitin chain (K63-polyUb) plays a key role in regulating RLRs signal pathway. Previous studies found that short chain K63-polyUb can effectively promote the oligomerization of RIG-ICARDs by covalent anchoring and non-covalent anchoring. The formed heterotetramer complexes (K63-polyUb-RIG-ICARDs) can activate MAVSCARD oligomerization and form the core of MAVS fibers. However, how K63-polyUb regulates the assembly of MDA5CARDs and the molecular mechanism of recruiting and activating MAVSCARD remains a scientific problem to be solved. Although K63-polyUb is required for MDA5 mediated MAVS activation, MDA5CARDs covalently modified by K63-polyUb have not been found. This implies the important role of unanchored K63-polyUb in MDA5-MAVS signal transmission.
In a study published in Immunity on October 13, 2021, ZHENG Jie's group of Shanghai Institute of Materia Medica, Chinese Academy of Sciences revealed the molecular mechanism that long chain, unanchored K63-polyUb promotes MDA5-MAVS assembly program and signal transduction.
Firstly, through hydrogen deuterium exchange mass spectrometry (HDX-MS), researchers found that MDA5CARDs prefer to combine longer K63-polyUbn than RIG-ICARDs (MDA5: n ≥ 8; RIG-I: n ≥ 3), but not K48-polyUbn (n ≥ 10) (Fig. 1).
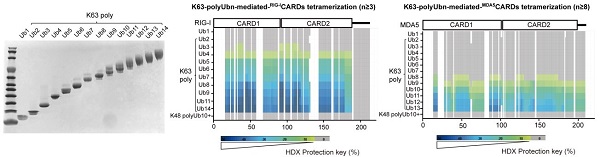
Figure 1. HDX-MS analysis of the effect of K63-polyUb (2 ≤ n ≤ 14) on the oligomerization of RLR CARDs
In order to study the assembly mechanism of oligomers of MDA5CARDs mediated by K63-polyUbn, the structure of the complex of MDA5CARDs-K63 polyUb13 with a resolution of 3.3 Angstrom was analyzed by Cryo-Electron Microscope (Cryo-EM) for the first time. The structure shows eight UB molecules surround a left-handed helical, cyclic CARDs tetramer. This is also the first Cryo-EM structure of MDA5CARDs with near atomic resolution. But how does the heterotetramer of MDA5CARDs-K63 polyUbn recruit its downstream signal protein MAVS? The researchers further obtained the "bottom-up" left-handed helical MDA5CARDs-MAVSCARD complex tethered by long chain K63-polyUb11 with a resolution of 3.2 Angstrom by Cryo-EM analysis.
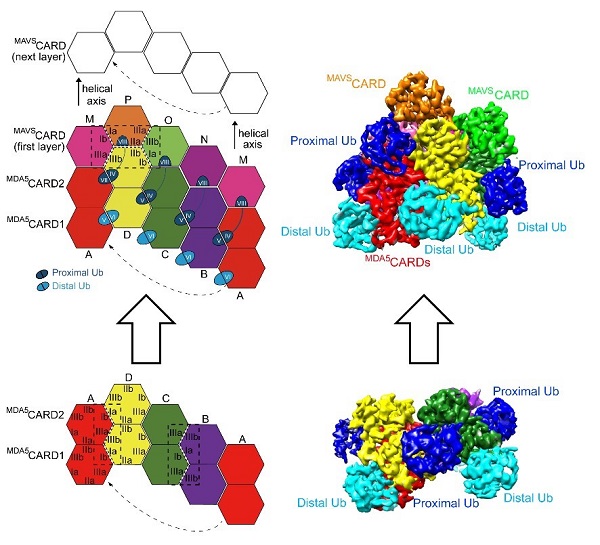
Figure 2. Single particle Cryo-EM shows the assembly mechanism of MDA5-MAVS (CARD-CARD) mediated by K63-polyUbn (n ≥ 8)
Moreover, the researchers proved for the first time that the CARDs of human MDA5 full-length protein are in an open conformation in the initial state and can bind to the long chain K63-polyUb10 through HDX-MS. However, in early studies, it was proved by HDX-MS that RIG-ICARDs has a closed conformation in the initial state. This also directly proved the great difference in the conformation of cards of RIG-I and MDA5 in solution. The researchers further found that the stability of K63-polyUb10 tethered MDA5CARDs complex in solution is regulated by the RNA dependent ATPase activity of MDA5, the effective recognition of RNA by MDA5 near mitochondrial membrane containing high concentration of ATP helps to remotely regulate the stability of K63 polyUb10-MDA5CARDs complex and help activate MAVS (Fig. 3).
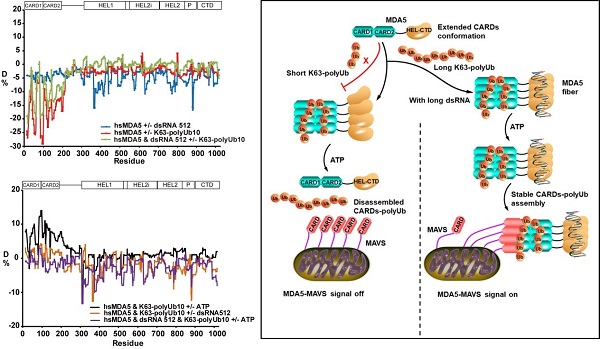
Figure 3. HDX-MS analysis of dynamic conformational changes and signal transduction mechanism of full-length MDA5 under the action of its recognition ligand or substrate (dsRNA / ATP / K63-polyUb)
This study analyzed the related structure of MDA5CARDs for the first time, and further revealed the immunological function of long chain, unanchored K63-polyUb as an endogenous agonist in cells, and provides a new clue for understanding the role of ubiquitin molecular diversity in anti-RNA virus natural immune signal transduction.
Article link: https://doi.org/10.1016/j.immuni.2021.09.008
Contact:
DIAO Wentong
Shanghai Institute of Materia Medica, Chinese Academy of Sciences
E-mail: diaowentong@simm.ac.cn




