Structural Basis for InhStructural Basis for Inhibiting HBV via HBx–Smc6 Interface: A New Therapeutic Directionibiting HBV via HBx–Smc6 Interface: A New Therapeutic Direction
Chronic hepatitis B virus (HBV) infection remains a major global health challenge, affecting approximately 250 million people worldwide, with about 80 million in China. Chronic infection is difficult to cure and often leads to cirrhosis and hepatocellular carcinoma (HCC). Globally, nearly one million people die each year from HBV-related cirrhosis and liver cancer, about half of them in China. Despite this burden, HBV has received relatively less attention from global health agendas and international pharmaceutical R&D, partly due to its lower incidence in Western countries. Current therapies (nucleos(t)ide analogues plus interferon) suppress viral replication but do not eliminate the virus, often requiring lifelong treatment and carrying a risk of drug resistance. Therefore, new mechanisms and therapeutic targets are urgently needed.
The HBV genome is ~3.2 kb and, through overlapping open reading frames, encodes surface proteins, core protein, polymerase, and the nonstructural protein X (HBx). After infection, the viral genome is converted in the nucleus into covalently closed circular DNA (cccDNA), which serves as a persistent viral reservoir and a key obstacle to cure. HBx is essential for multiple steps of the viral life cycle and is also considered a key oncoprotein. The Smc5/6 complex has been identified as a host restriction factor that suppresses cccDNA transcription early in infection. To counteract this, HBx hijacks the host protein DDB1 and recruits the CRL4 ubiquitin ligase to degrade Smc5/6, thereby relieving transcriptional repression. Because HBx is intrinsically disordered and highly flexible, its full-length structure and the precise mechanism of Smc5/6 recognition have remained elusive – even AlphaFold 3 did not yield high-confidence models, limiting drug development.
In a study published in Cell Research, a collaborative team of researchers led by WANG Lanfeng from the Shanghai Institute of Materia Medica, Chinese Academy of Sciences, YUAN Zhenghong from Fudan University, and RAO Zihe from Tsinghua University, expressed and purified a ten-subunit assembly containing HBx, CRL4, and the human Smc5/6 complex using a baculovirus–insect cell system, and determined its three-dimensional structure by single-particle cryo‑electron microscopy.
For the first time, the structure of near-full-length HBx was resolved, revealing a Y-shaped conformation stabilized by zinc ions – markedly different from AlphaFold 3 predictions. The “lower arm” of HBx binds tightly to DDB1, while the “upper arm” uses a composite helix-turn-helix (HTH) hydrophobic pocket to precisely recognize a “leucine key” on Smc6. Disrupting this interface completely abolishes HBx-mediated ubiquitination and degradation of Smc6, and inhibits HBV transcription and replication. This key–lock interaction is the molecular basis for HBx‑dependent Smc5/6 degradation.
The Smc5/6 complex also maintains genomic stability and participates in DNA damage repair. In hepadnavirus-infected mammals, HBx‑mediated degradation of Smc5/6 compromises genomic stability. Moreover, in chronically infected patients, nearly 100% of infected hepatocytes carry integrated HBV DNA. These integrated viral genes continuously express various HBx truncation mutants, and those retaining the composite HTH pocket can still degrade Smc5/6. Sequence alignment shows that the residues forming this pocket are fully conserved across all HBV genotypes and even in ancient HBV strains (thousands of years old). Evolutionary analysis indicates that the X gene emerged in mammalian hepatitis viruses and that this hydrophobic pocket is highly conserved. On the host side, the “leucine key” on Smc6 appeared as early as in birds and remains fully conserved in mammals, suggesting co‑evolution. This aligns with the fact that HCC occurs only in infected mammals, implying that HBx‑mediated Smc5/6 degradation may significantly contribute to hepatocarcinogenesis. Therefore, blocking the HBx–Smc6 interaction could both inhibit viral replication and delay progression from hepatitis to HCC.
Using the conserved composite HTH pocket as a target, the team performed high‑throughput virtual screening of an FDA‑approved compound library and identified the small molecule Tranilast. This compound competitively binds the HTH pocket, blocking the HBx–Smc6 interaction and thereby inhibiting Smc6 ubiquitination and degradation. Both in vitro and cell‑based assays showed dose‑dependent inhibition, providing proof‑of‑concept for targeting this interface. Notably, this approach targets both cccDNA‑derived and integration‑derived HBx, and could both suppress viral replication and potentially delay hepatocarcinogenesis. This work opens a new direction for hepatitis B therapy.
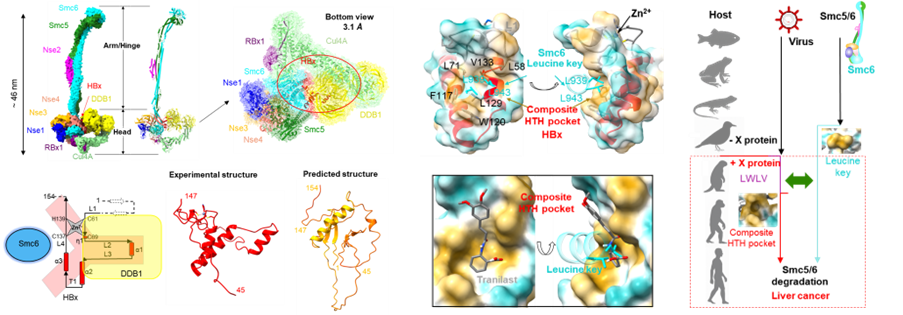
Structural determination of the HBx–CRL4–Smc5/6 ten-subunit complex, high-resolution three-dimensional structure of HBx, and proof-of-concept for a drug targeting its constitutive HTH pocket (Image by WANG Lanfeng’s lab)
DOI: 10.1038/s41422-026-01251-7
Link:https://www.nature.com/articles/s41422-026-01251-7
Keywords: hepatitis B virus, HBx, Smc5/6 complex, cryo-EM
Contact:
DIAO Wentong
Shanghai Institute of Materia Medica
E-mail: diaowentong@simm.ac.cn




